Quantitative Analysis of the Chemotactic Motility of Amoeboid Cells
Grant Number: 1 R01 GM084227-01
Funding Agency: NIH. (National Institute for General Medical Sciences).
Lead PI Professor Juan C. Lasheras with Professor Richard Firtel (Cell and Developmental Biology) (co-PI) and Professor Juan C. Del Alamo (co-PI).
Researchers
- Baldomero Alonso-Latorre
- Juan C. del Álamo
- Ruedi Meili (Firtel’s lab)
- Effie Bastounis
Eukaryotic cell motility is essential for many physiological processes such as embryonic development, tissue renewal, and the function of the immune system. The amoeba Dictyostelium discoideum has proven to be an excellent model for the chemotactic migration of amoeboid cells such as leukocytes. These cells move in response to external stimuli by remodeling their cytoskeleton and their substrate adhesions. This process is controlled by a complex network of signaling pathways which drive a simple repetitive sequence of mechanical actions coordinated in space and time (motility cycle). The mechanisms behind this biophysical integration are still poorly understood. Preliminary studies in my group (publication A-I-72) suggest that the motility cycle follows a simple hyperbolic law (VT= constant =L) over a wide range of conditions for several mutants tested (V=speed, T=period of the mechanical energy of cell adhesion, L= constant step length). Mechanical processes alone cannot explain this hyperbolic law because amoeboid cells do not keep a fixed size during migration unlike, for example, an inchworm, which suggests an integrated biophysical process as a possible explanation. We therefore hypothesize that the amoeboid motility cycle is governed by a step of constant length, L, which is determined by multiple interconnected processes operating during different stages of the cycle.
The first aim of our research is to determine the dependence of L and T on external variables such as the shape and magnitude of the chemoattractant gradient and the stiffness of the substrate. The second aim is to study the molecular basis for L and T by studying the role of candidate molecules with known or suspected involvement in the different stages of the cycle. For this purpose we are using knock out and transgenic cell lines as well as pharmacological inhibitors. We are also investigating actin polymerization and its regulation, a process likely involved in the determination of L. We are simultaneously measuring the spatial and temporal changes in the distribution of fluorescently tagged signaling or cytoskeletal proteins and the evolution of the traction forces that mediate each stage of the cell motility cycle, while also monitoring the changes in the geometry and the cytoplasm rheology of the cell all the time. Conditional and phase statistics enable us to integrate all the data and build a numerical representation of a canonical cell undergoing a canonical motility cycle. These combined measurements provide the quantitative information needed to connect specific biochemical processes to each of the physical events in the motility cycle by statistical correlation. We are working on models to predict the motility behavior of cells and our analysis is expected to be applicable to cells with similar motility properties like leukocytes. Future therapeutic approaches to treat diseases involving motile cells such as cancer, destructive inflammation, and osteoporosis will benefit from such a model.
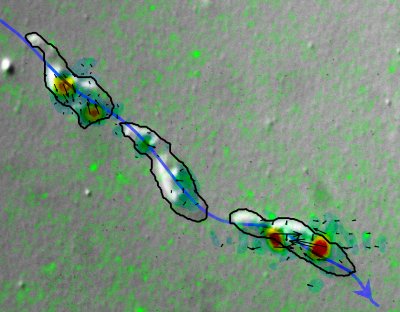
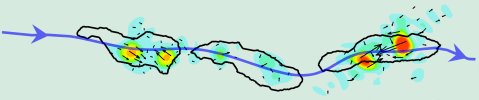
Composite image of a Dictyostelium discoideum cell crawling on a elastic substrate seeded with green fluorescent marker beads at the times t=0s, t= 100s and t= 200s. The solid black contours show the cells' outline at these time points, and the blue curve indicates the trajectory which the cell follows up the chemoattractant gradient. The magnitude of the traction forces (black arrows) and the stresses (color contours) exerted by the cell at each instant was calculated from the displacement of the marker beads using our traction force microscopy method.
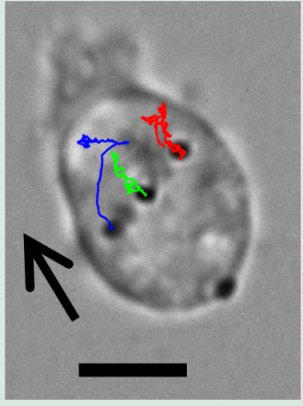
Bright field image of a moving Dictyostelium cell that has been injected with 0.5-micron gold microbeads (black dots). The curves show the trajectories of three microbeads tracked during 10 seconds. The cell is moving in the direction pointed by the arrow. The scale bar is 5 microns long. This experiment was performed in collaboration with Alex Groisman and Orit Shefi (Macagno Lab).
Videos of Research
Related Papers
![]() J.C. Del Álamo, R. Meili, B.Alonso-Latorre, J. Rodríguez-Rodríguez, A. Aliseda, R. Firtel and J.C. Lasheras. "Spatio-temporal Analysis of Eukaryotic Cell Motility by Improved Force Cytometry." Proc. Nat. Acad. Sciences of the USA. vol. 104 no. 33 pp. 13211-13212, (2007).
J.C. Del Álamo, R. Meili, B.Alonso-Latorre, J. Rodríguez-Rodríguez, A. Aliseda, R. Firtel and J.C. Lasheras. "Spatio-temporal Analysis of Eukaryotic Cell Motility by Improved Force Cytometry." Proc. Nat. Acad. Sciences of the USA. vol. 104 no. 33 pp. 13211-13212, (2007).
![]() B. Alonso-Latorre, R. Meili, E. Bastounis, J.C. Del Álamo, R. Firtel and J.C. Lasheras. Distribution of traction forces associated with shape changes during amoeboid cell migration. IEEE Eng. Med Biol. Soc. 2009:3346-9. (2009).
B. Alonso-Latorre, R. Meili, E. Bastounis, J.C. Del Álamo, R. Firtel and J.C. Lasheras. Distribution of traction forces associated with shape changes during amoeboid cell migration. IEEE Eng. Med Biol. Soc. 2009:3346-9. (2009).
![]() R. Meili, B. Alonso-Latorre, J.C. Del Álamo, R.A. Firtel and J.C. Lasheras. "Myosin II is Essential for the Spatial Organization of Traction Forces during Cell Motility." Molecular Biology of the Cell. Vol. 21 (3) pp. 405-417, (2010).
R. Meili, B. Alonso-Latorre, J.C. Del Álamo, R.A. Firtel and J.C. Lasheras. "Myosin II is Essential for the Spatial Organization of Traction Forces during Cell Motility." Molecular Biology of the Cell. Vol. 21 (3) pp. 405-417, (2010).
![]() J.C. Lasheras, R. Meili, B. Alonso-Latorre, J.C. Del Álamo, R.A. Firtel. "Distribution of Traction Forces and Intracellular Markers Associated with Shape Changes During Amoeboid Cell Migration." Int. Journal of Transport Phenomena. (2011).
J.C. Lasheras, R. Meili, B. Alonso-Latorre, J.C. Del Álamo, R.A. Firtel. "Distribution of Traction Forces and Intracellular Markers Associated with Shape Changes During Amoeboid Cell Migration." Int. Journal of Transport Phenomena. (2011).
![]() E. Bastounis, R. Meili, B. Alonso-Latorre, J.C. Del Álamo, J.C. Lasheras and R.A. Firtel. "Role of the Scar/WAVE Complex in Regulating Traction Forces During Amoeboid Motility." Molecular Biology of the Cell. Vol. 22 (21) pp: 3995-4003. (2011).
E. Bastounis, R. Meili, B. Alonso-Latorre, J.C. Del Álamo, J.C. Lasheras and R.A. Firtel. "Role of the Scar/WAVE Complex in Regulating Traction Forces During Amoeboid Motility." Molecular Biology of the Cell. Vol. 22 (21) pp: 3995-4003. (2011).
![]() B. Alonso-Latorre, J.C. Del Álamo, R. Meili, R.A. Firtel and J.C. Lasheras. "Strain energy modes in migrating amoeboid cells." J. Cellular and Molecular Bioengineering. Vol. 4, Number 4, pp. 603-615 (2011).
B. Alonso-Latorre, J.C. Del Álamo, R. Meili, R.A. Firtel and J.C. Lasheras. "Strain energy modes in migrating amoeboid cells." J. Cellular and Molecular Bioengineering. Vol. 4, Number 4, pp. 603-615 (2011).
J.C. Del Álamo, B. Álvarez, R. Meili, R.A. Firtel and J.C. Lasheras. "Three-Dimensional Quantification of Cellular Traction Forces and Mechanosensing of Thin Substrata by Fourier Traction Force Microscopy." Submitted. PLOS ONE. (2012).
E. Bastounis, R. Meili, J.C. Del Álamo, R.A. Firtel and J.C. Lasheras. "Chemotaxing amoeboid cells migrate by switching between distinct adhesion dynamics and contractility modes." Developmental Cell. Submitted. (2012).

